DPLM-2: A Multimodal Diffusion Protein Language Model
What is DPLM-2: A Multimodal Diffusion Protein Language Model
DPLM-2 is a protein AI model that can work with both sequence and structure at the same time. It can write protein sequences and also predict 3D shapes, all in one go.
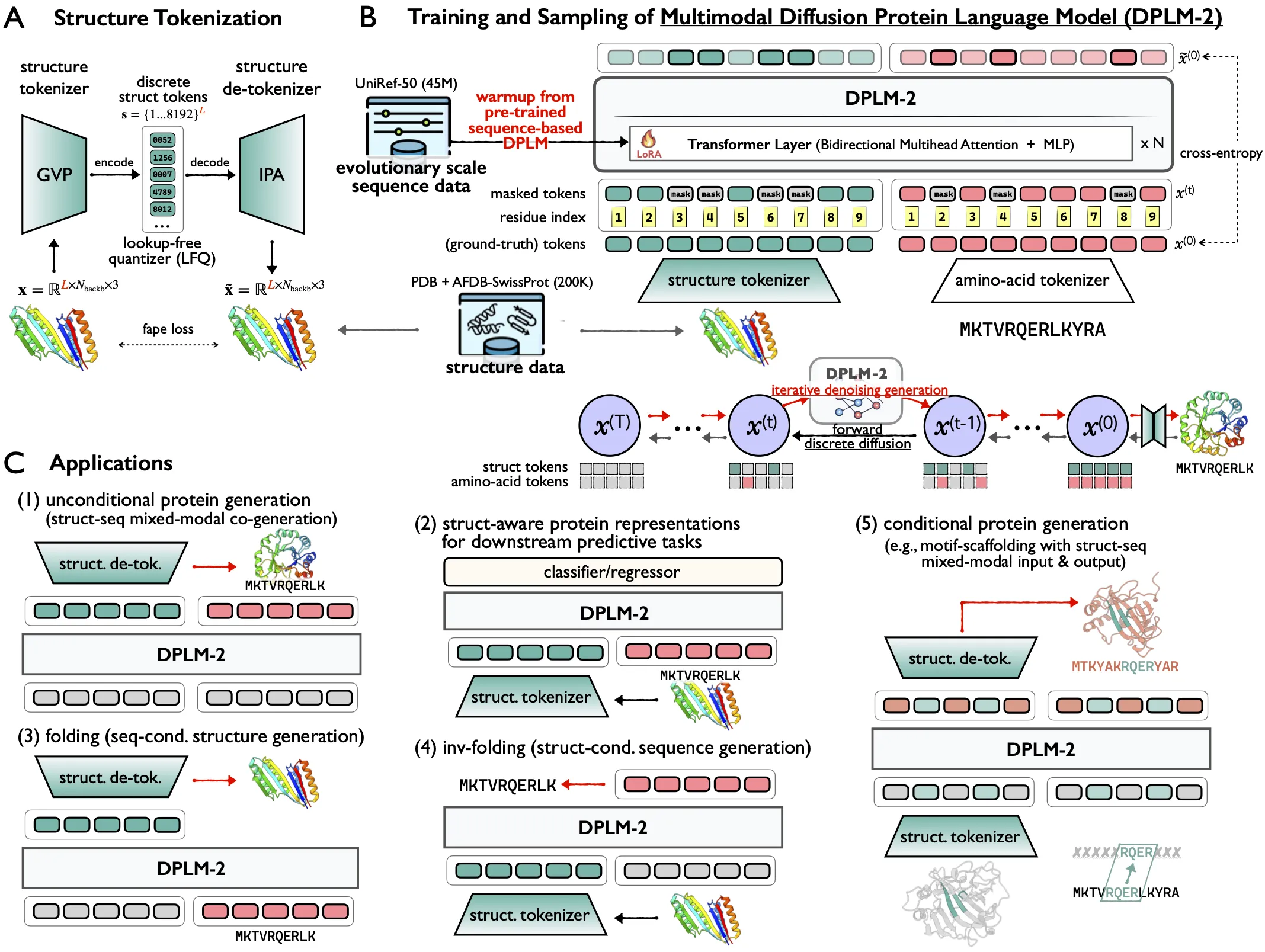
This helps researchers design proteins faster. It also supports tasks like folding (structure from sequence), inverse folding (sequence from structure), and motif scaffolding (keep a key part, design the rest).
DPLM-2: A Multimodal Diffusion Protein Language Model Overview
DPLM-2 builds on the DPLM family and adds structure modeling to the same model that learns sequences. Instead of using one model for sequence and another for structure, it treats both together.
Here is a quick view of the project.
| Item | Details |
|---|---|
| Type | Multimodal protein language model (sequence + structure) |
| Purpose | Co-generate protein sequences and 3D structures in one step |
| Core Idea | Discrete diffusion model with a tokenizer that turns 3D coordinates into tokens |
| Key Tasks | Co-generation, folding, inverse folding, motif scaffolding, representation learning |
| Extra Variant | DPLM-2 Bit (bit-based structure tokens for finer control) |
| Pretrained Models | airkingbd/dplm_650m, airkingbd/dplm2_650m, airkingbd/dplm2_bit_650m |
| Training Data (DPLM-2) | PDB (experimental) + SwissProt with AF2-predicted structures |
| Codebase | ByProt-based implementation |
| Source | Project site and GitHub (training + inference + scripts) |
| Best For | Protein design teams, biology students, and ML+bio labs |
| Project Page | https://bytedance.github.io/dplm/dplm-2 |
You can also learn more about the team background here: ByteDance overview.
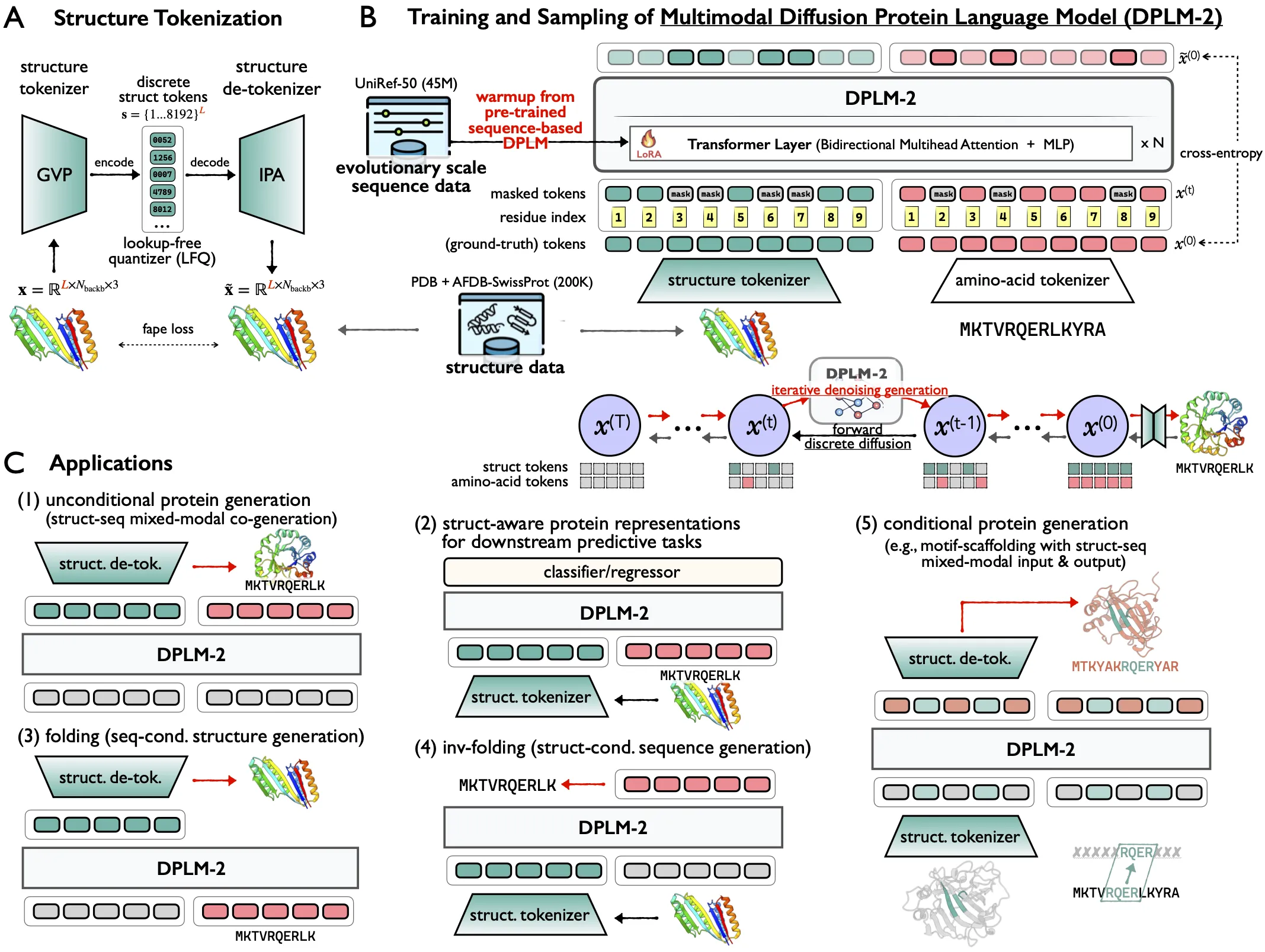
DPLM-2: A Multimodal Diffusion Protein Language Model Key Features
- Sequence–structure co-generation: Create a protein sequence and its 3D structure together.
- Folding from sequence: Predict a structure from an input sequence, with strong results close to tools like ESMFold.
- Inverse folding: Given a target backbone, generate sequences that are likely to fold to it.
- Motif scaffolding: Keep important motifs and design the rest around them with multimodal inputs.
- Representation learning: Get structure-aware embeddings for downstream tasks.
- Controllable options: Add simple controls such as secondary structure hints.
DPLM-2: A Multimodal Diffusion Protein Language Model Use Cases
- Design new proteins with both sequence and structure in one run.
- Predict structures for candidate sequences during screening.
- Redesign sequences to match a known backbone shape.
- Build scaffolds that preserve a functional motif.
- Train simple predictors on top of learned embeddings for property tasks.
How It Works
Proteins have letters (amino acids) and shapes (3D coordinates). DPLM-2 turns both into tokens and learns how they relate.
It uses diffusion to iteratively refine tokens and produce clean outputs. A warm-up method starts from a strong sequence-only model (DPLM), then teaches the model about structure without losing past knowledge.
DPLM-2 Bit improves structure prediction by predicting bits of the structure code, not just a single index. This makes structure modeling more fine-grained. For more friendly AI explainers, see our hub: learn more topics.
Getting Started: Installation & Setup
Follow these steps exactly as shown.
# clone project
git clone --recursive https://url/to/this/repo/dplm.git
cd dplm
# create conda virtual environment
env_name=dplm
conda create -n ${env_name} python=3.9 pip
conda activate ${env_name}
# automatically install everything else
bash scripts/install.sh
Load Pretrained Models
Use these lines to load checkpoints in Python.
from byprot.models.dplm import DiffusionProteinLanguageModel as DPLM
from byprot.models.dplm2 import MultimodalDiffusionProteinLanguageModel as DPLM2
from byprot.models.dplm2 import DPLM2Bit
dplm = DPLM.from_pretrained("airkingbd/dplm_650m").cuda()
dplm2 = DPLM2.from_pretrained("airkingbd/dplm2_650m").cuda()
dplm2_bit = DPLM2Bit.from_pretrained("airkingbd/dplm2_bit_650m").cuda()
Quick Examples: Generate Proteins
Here are ready-to-run examples.
Protein sequence generation:
from generate_dplm import initialize_generation
input_tokens = initialize_generation(
length=200,
num_seqs=5,
tokenizer=dplm.tokenizer,
device=next(dplm.parameters()).device
)
samples = dplm.generate(
input_tokens=input_tokens,
max_iter=500,
)
print([''.join(seq.split(' ')) for seq in dplm.tokenizer.batch_decode(samples, skip_special_tokens=True)])
Protein sequence-structure co-generation:
User can check the generated sequence and structure in the ./generation-results folder.
from generate_dplm2 import initialize_generation, save_results
input_tokens = initialize_generation(
task="co_generation",
length=200,
num_seqs=5,
tokenizer=dplm2.tokenizer,
device=next(dplm2.parameters()).device
)[0]
samples = dplm2.generate(
input_tokens=input_tokens,
max_iter=500,
)
save_results(
outputs=samples,
task="co_generation",
save_dir="./generation-results/dplm2_generation",
tokenizer=dplm2.tokenizer,
struct_tokenizer=dplm2.struct_tokenizer, save_pdb=True
)
samples = dplm2_bit.generate(
input_tokens=input_tokens,
max_iter=500,
)
save_results(
outputs=samples,
task="co_generation",
save_dir="./generation-results/dplm2_bit_generation",
tokenizer=dplm2_bit.tokenizer,
struct_tokenizer=dplm2_bit.struct_tokenizer
)
Training Guide
Download datasets for faster local loading.
DPLM (UniRef50):
bash scripts/download_uniref50_hf.sh
DPLM-2 (PDB + SwissProt with AF2 structures):
bash scripts/download_pdb_swissprot.sh
Train DPLM (sequence-only). This runs on one node with 8 A100 GPUs:
export CUDA_VISIBLE_DEVICES=0,1,2,3,4,5,6,7
max_tokens=8192
accumulate_grad_batches=16
# this means the effective batch size is #GPUs(8) * max_tokens(8192) * accumulate_grad_batches(16), resulting in approximately 1 million.
exp=dplm/dplm_650m
model_name=dplm_650m
python train.py \
experiment=${exp} name=${model_name} \
datamodule.max_tokens=${max_tokens} \
trainer.accumulate_grad_batches=${accumulate_grad_batches}
Train DPLM-2 (sequence + structure) with warm-up and LoRA:
export CUDA_VISIBLE_DEVICES=0,1,2,3,4,5,6,7
max_tokens=8192
accumulate_grad_batches=1
# this means the effective batch size is #GPUs(8) * max_tokens(8192) * accumulate_grad_batches(1), resulting in approximately 64 thousand.
exp=dplm2/dplm2_650m
model_name=dplm2_650m
python train.py \
experiment=${exp} name=${model_name} \
datamodule.max_tokens=${max_tokens} \
trainer.accumulate_grad_batches=${accumulate_grad_batches}
Train DPLM-2 Bit (bit-based structure tokens):
export CUDA_VISIBLE_DEVICES=0,1,2,3,4,5,6,7
max_tokens=8192
accumulate_grad_batches=1
# this means the effective batch size is #GPU(8) * max_tokens(8192) * accumulate_grad_batches(1), resulting in approximately 64 thousand.
exp=dplm2/dplm2_bit_650m
model_name=dplm2_bit_650m
python train.py \
experiment=${exp} name=${model_name} \
datamodule.max_tokens=${max_tokens} \
trainer.accumulate_grad_batches=${accumulate_grad_batches}
Need a quick profile of our work? Check our story here: learn about us.
Unconditional Generation from CLI
Generate sequences with a pre-trained DPLM model and score with ESMFold.
model_name=dplm_650m # choose from dplm_150m, dplm_650m, dplm_3b
output_dir=generation-results/${model_name}/uncond_generation
mkdir -p generation-results
python generate_dplm.py --model_name airkingbd/${model_name} \
--seq_lens 100 200 300 400 500 \
--saveto ${output_dir}
# Evaluation
bash anylasis/plddt_calculate.sh ${output_dir} # compute pLDDT using ESMFold
Performance & Showcases
DPLM-2 co-generates sequences and structures in a single step, removing the need for a two-stage run. It shows strong results across folding, inverse folding, and motif scaffolding.
In folding from sequence, reported results are close to strong baselines like ESMFold. In inverse folding, it produces sequences that are likely to fit the target backbone.
DPLM-2 Bit further improves structure modeling by predicting bits of the structure code, which helps with fine details in shape. Sampling defaults were later tuned to “annealing@2.0:0.1” for better results.
Tips and Notes
- Hardware: examples show 8x A100 GPUs. Adjust max_tokens and accumulate_grad_batches to keep total tokens per step in the same range.
- Data: use the provided download scripts to cache datasets locally. This speeds up training.
- Results: co-generation scripts can save PDB files to the generation-results folder for quick checks.
FAQ
What does “multimodal” mean here?
It means the model reads and writes both protein letters (sequence) and 3D points (structure). DPLM-2 learns how the two connect.
Do I need structure data to start?
For DPLM-2 training, yes. Use the PDB and SwissProt with AF2 structure dataset via the provided download script. For sequence-only work, DPLM is enough.
How big are the models?
The repo mentions DPLM sizes like 150M, 650M, and 3B. DPLM-2 examples use a 650M model name in scripts.
What files do the generation scripts save?
They write sequences and, for DPLM-2, structures to the generation-results folder. You can also ask the script to save PDB files.
Where can I learn more about the company or related topics?
You can read a quick overview here: company background. For guides and news, see our main site.
Image source: DPLM-2: A Multimodal Diffusion Protein Language Model